disimrank
Gene target ranking by driven input single input motif Matlab code.
MATLAB Gaussian Process Transcrption Factor Target Ranking Toolbox
This page contains a MATLAB implementation of the transcription factor target ranking methodology based on Gaussian process differential equation models.
An R implementation of the same methods is available from the authors upon request.
Author information
This package was created by Antti Honkela.
Release Information
Current release is 0.11.
As well as downloading the DISIMRANK software you need to obtain the toolboxes specified below. These can be downloaded using the same password you get from registering for the DISIMRANK software.
| Toolbox | Version |
|---|---|
| NETLAB | 3.3 |
| OPTIMI | 0.132 |
| NDLUTIL | 0.161 |
| MLTOOLS | 0.134 |
| KERN | 0.225 |
| PRIOR | 0.22 |
| GPSIM | 0.1211 |
Minor fix to demPlotMef2Models.m.
Version 0.1
This version includes scripts used to run the experiments in the PNAS paper.
Data
The PNAS experiments depend on a Drosophila data set (35 MB), which includes a pre-processed version of the developmental expression time series data of Tomancak et al. as well as various validation data sets. We thank Dr. Tomancak for the kind permission to redistribute the data.
Examples
In order to run the demos, you need the above Drosophila data set. The command drosLoadData (also used internally by many of the scripts) will attempt to load it from the current working directory and its subdirectory data.
Models can be fitted for individual genes and the results visualised using the command
>> demPlotModels
>>This will create independent visualisations of model for the three repeated experiments, the last of which are:
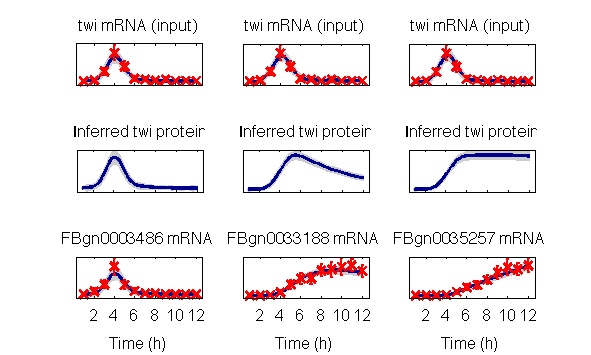
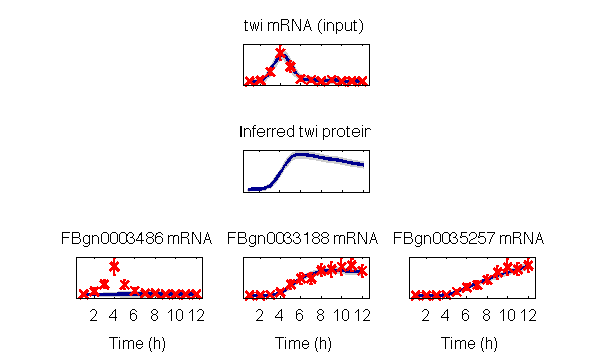
The package can also be used to rerun the rankings reported in the paper. This is a time-consiming process that may take several days in serial execution. If desired, these may nevertheless be run using the command
>> demRunRankings
>>Page updated on Thu Apr 15 12:13:57 2010